https://enanomapper.adma.ai
-
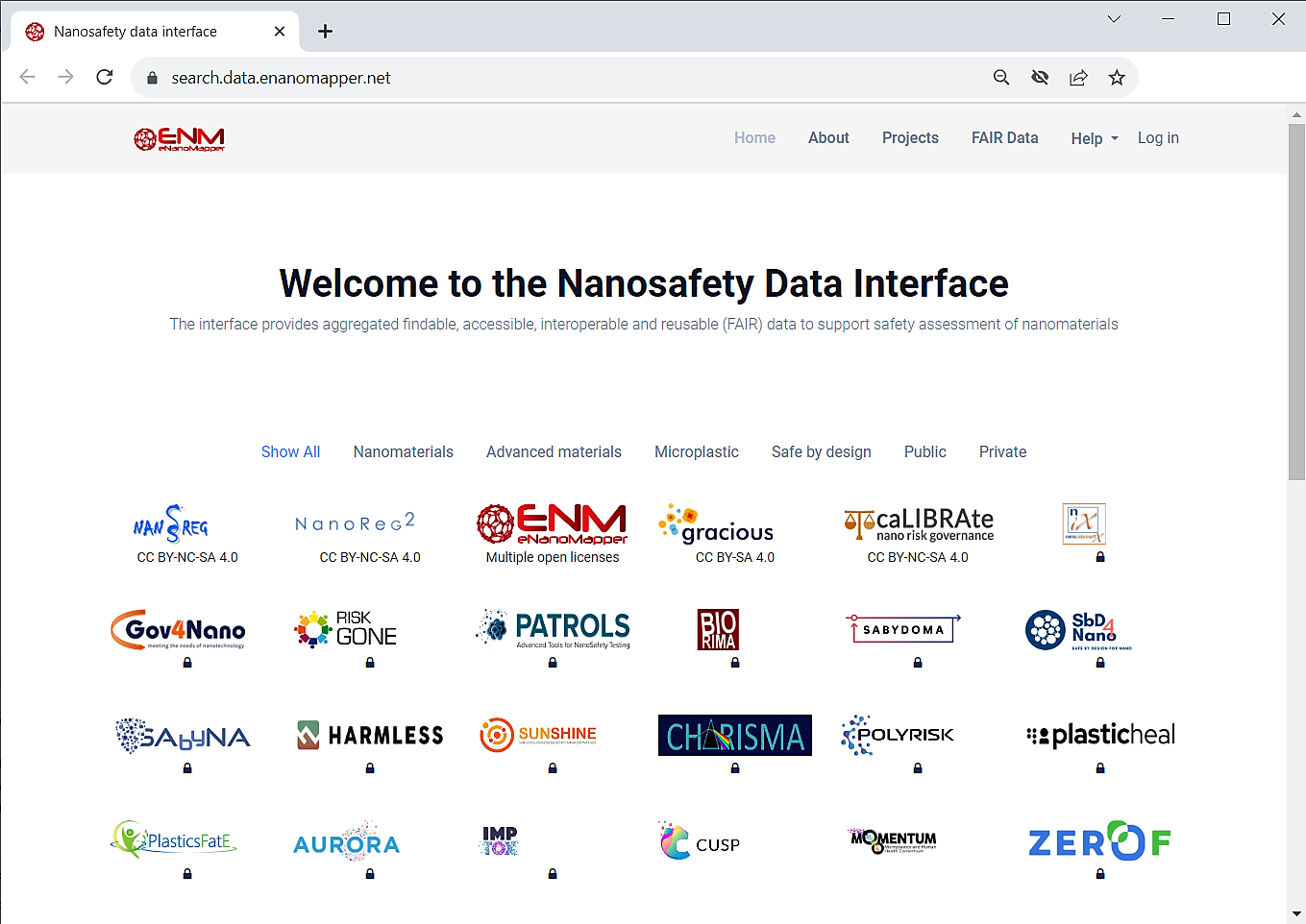
The Gracious database is publicly accessible.
-
Click on GRACIOUS icon to go to the H2020 GRACIOUS database main project page
-
Click on GRACIOUS icon to go to the H2020 GRACIOUS database main project page
Main project page
On the main page there are several links to the Search, Template Wizard, User guide (this page), etc.
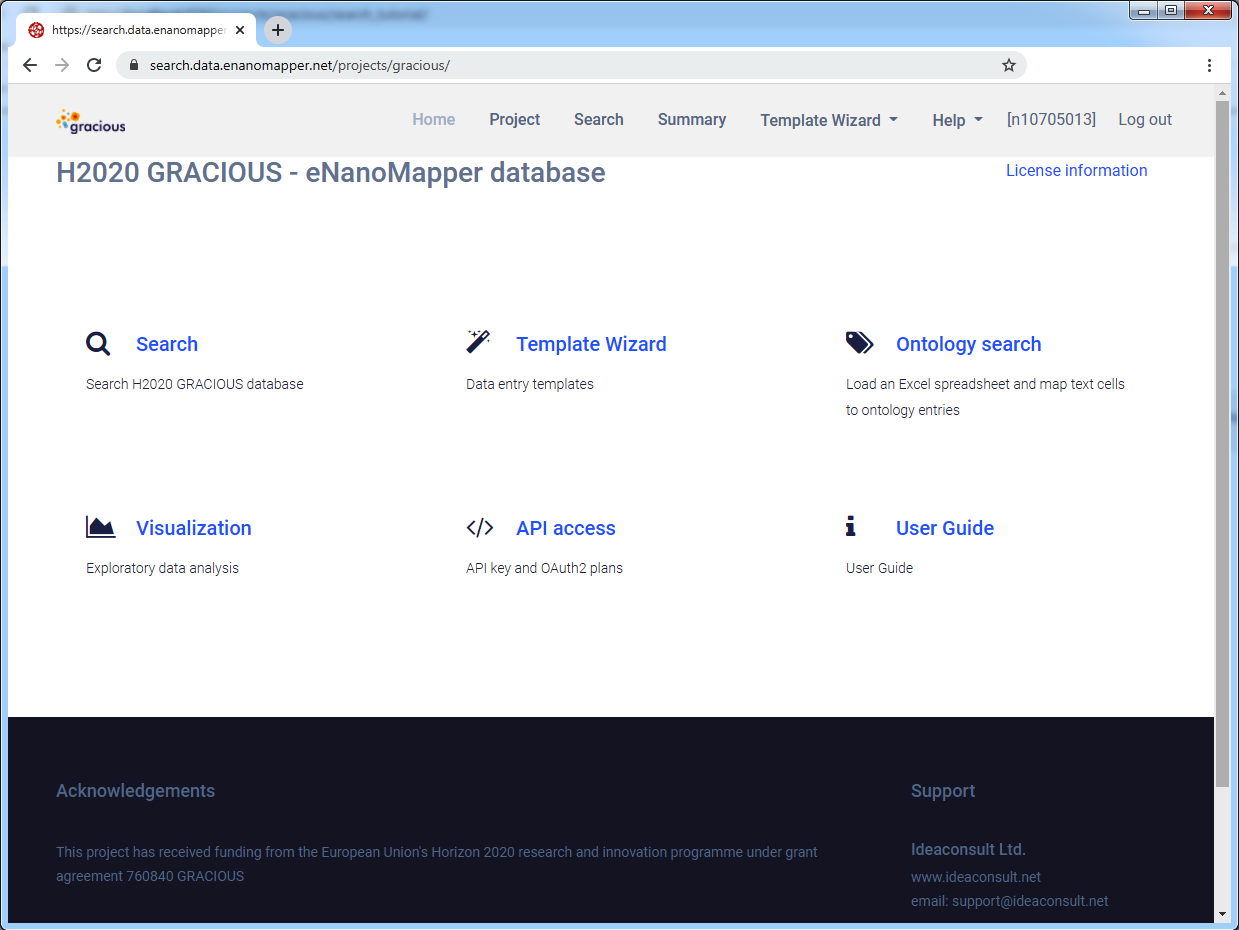
Click on the Search link to go to the Search page
Search page
-
On the search page there is a search box on the top right , a summary panel on the left and the results are shown at the main panel.
-
The database includes results from multiple sources.
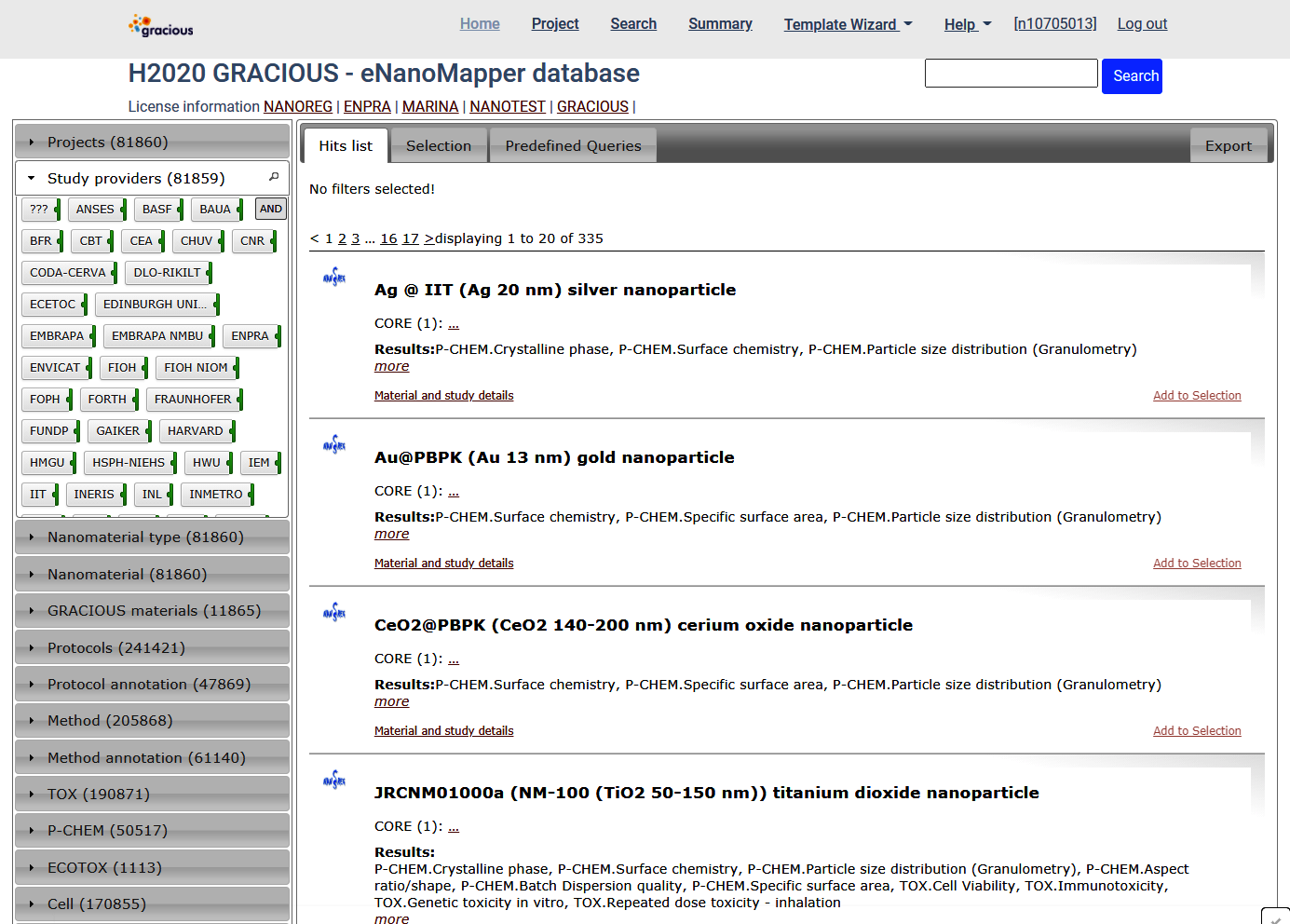
There are several summaries at the left panel:
- Project
- Study providers
- Nanomaterial type
- Nanomaterials
- Protocols
- Protocol annotation
- Method
- Method annotation
- P-Chem
- Tox
- Ecotox
- Cell
- Experiment year
- References
- Endpoint
- Endpoint annotation
- Release
Every panel is expandable and shows the types of elements found for the particular query
Expand the Projects panel and click on GRACIOUS. This will filter the results and only data generated by GRACIOUS project will be shown.
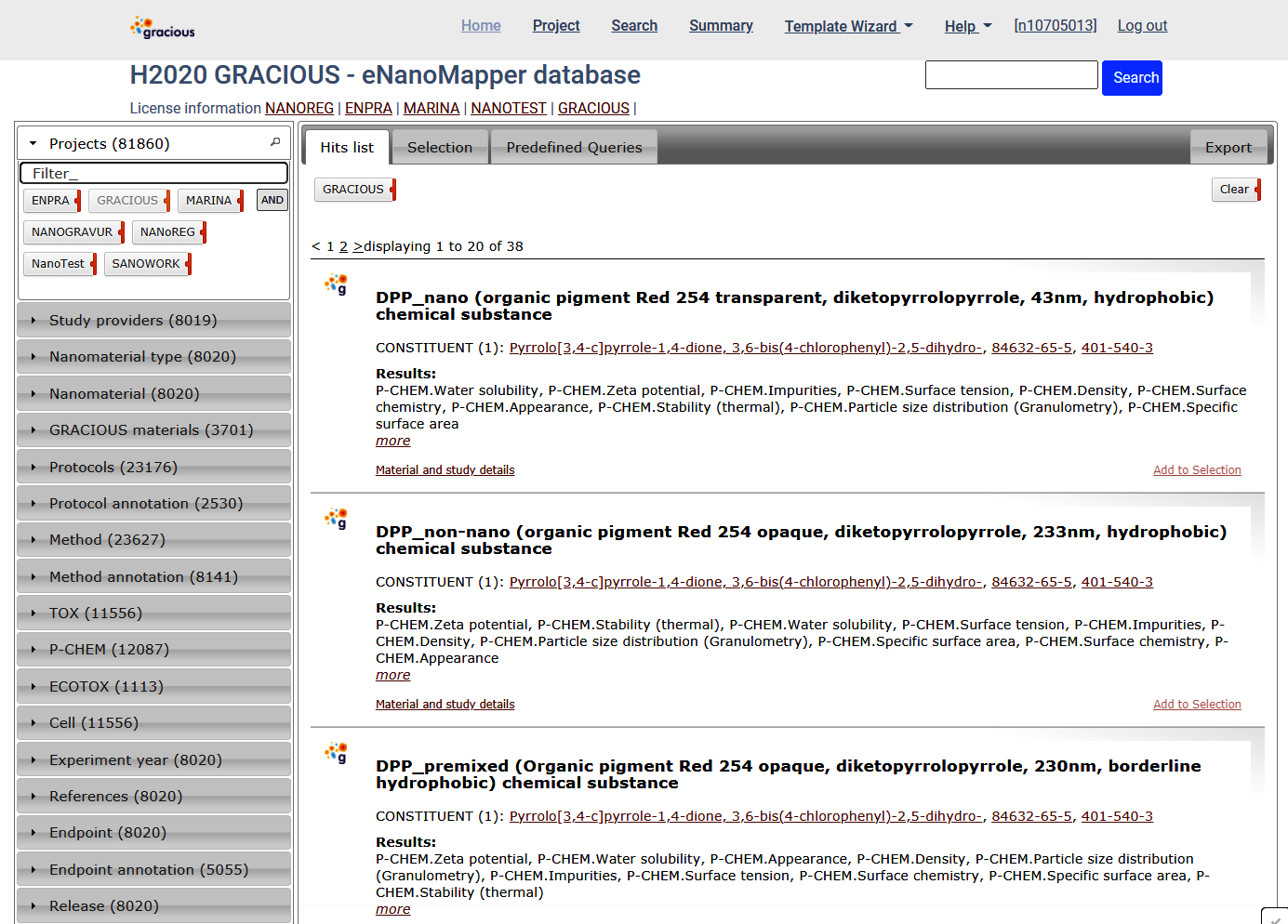
The filters can be combined.
Expand the Nanomaterial type panel and select quantum dot. These are ontology terms assigned to each material , e.g. NPO_589
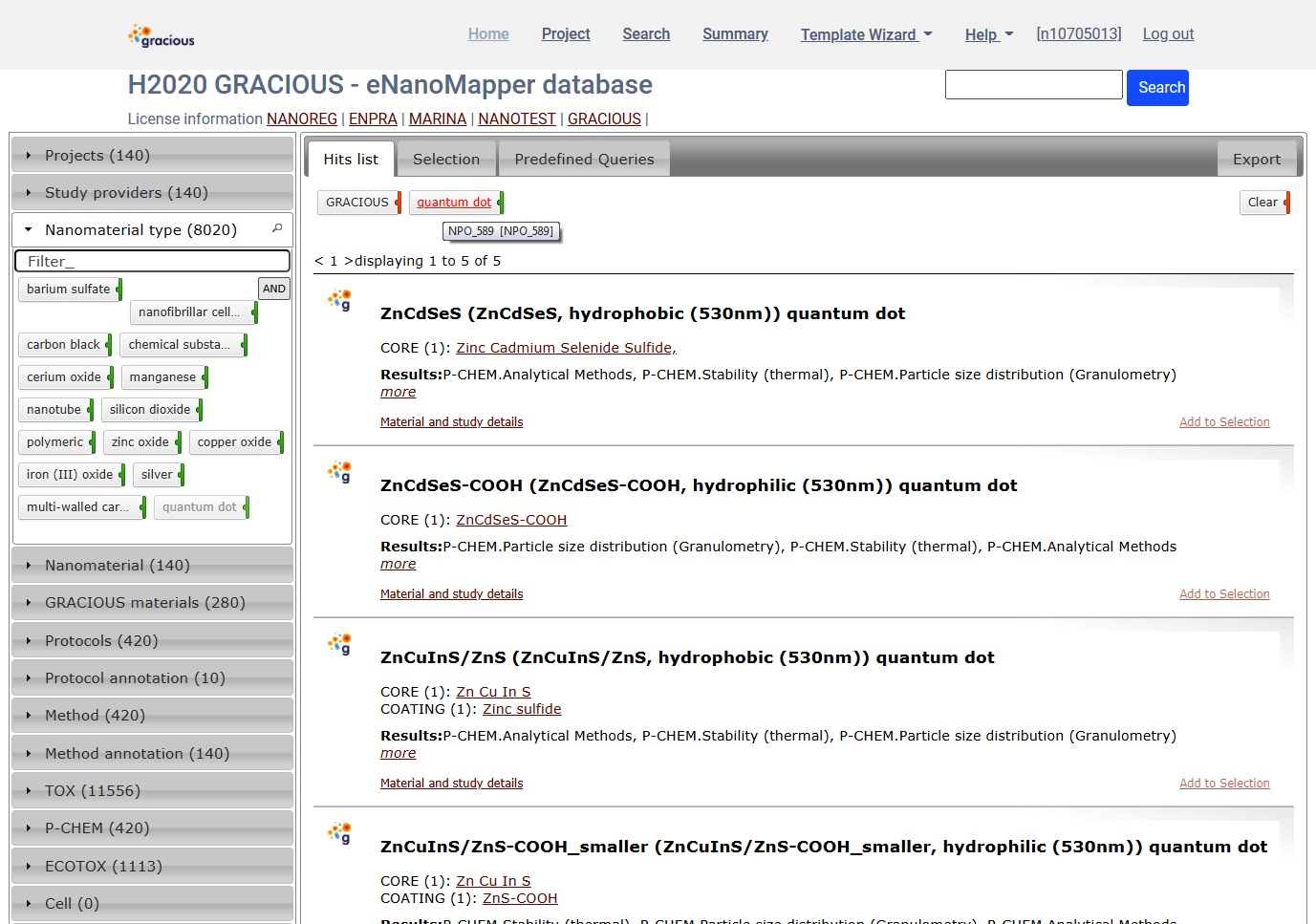
The panels show the types and counts of elements found for the particular query
i.e, the GRACIOUS project and quantum dot materials in this case.
The Methods panel shown indicate there is GRACIOUS data for quantum dot measured by these methods. The Method annotation panel contains the ontology terms denoting the same methods.
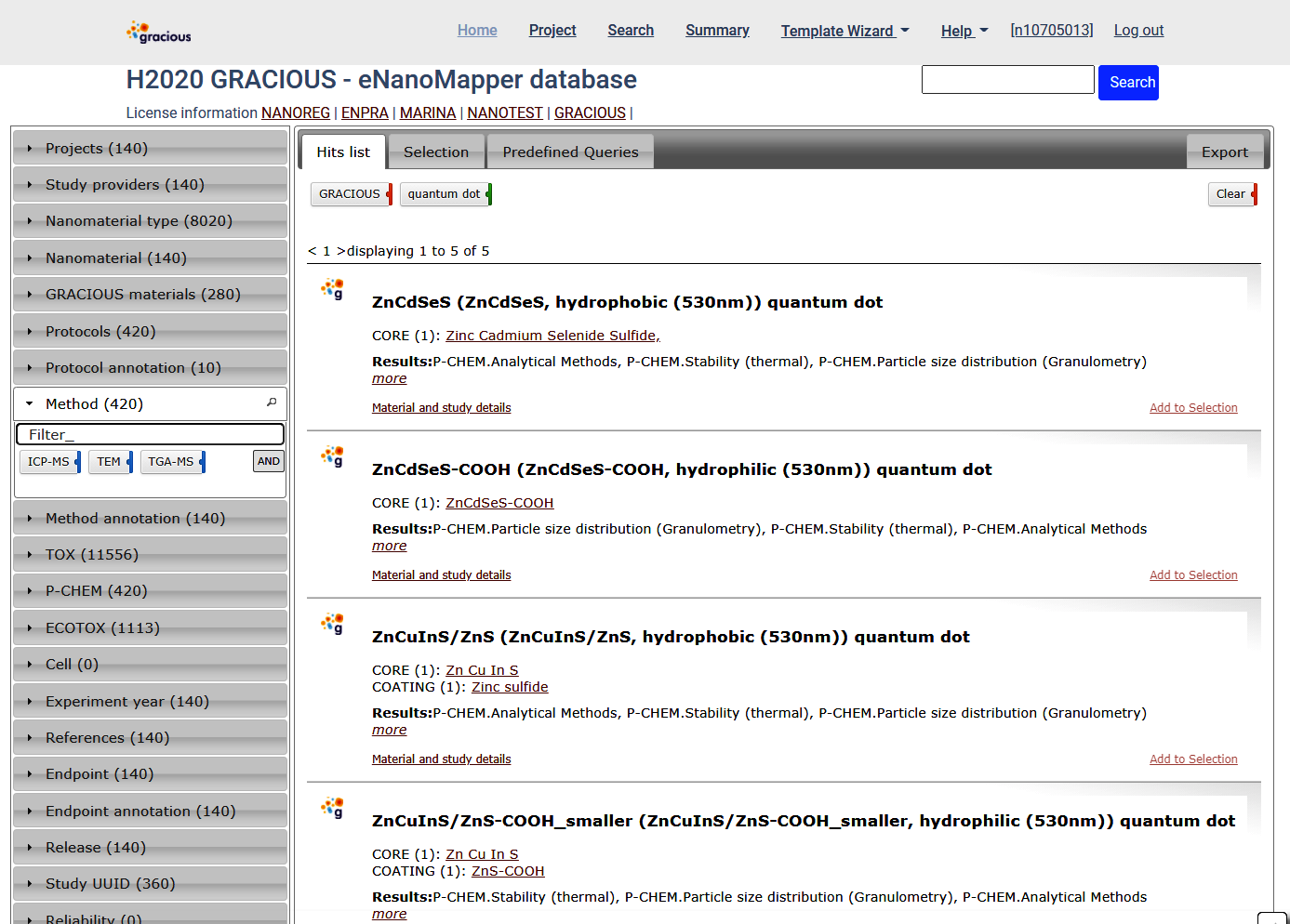
Exploring the data
Per material - click the Material and Study details link for particular material
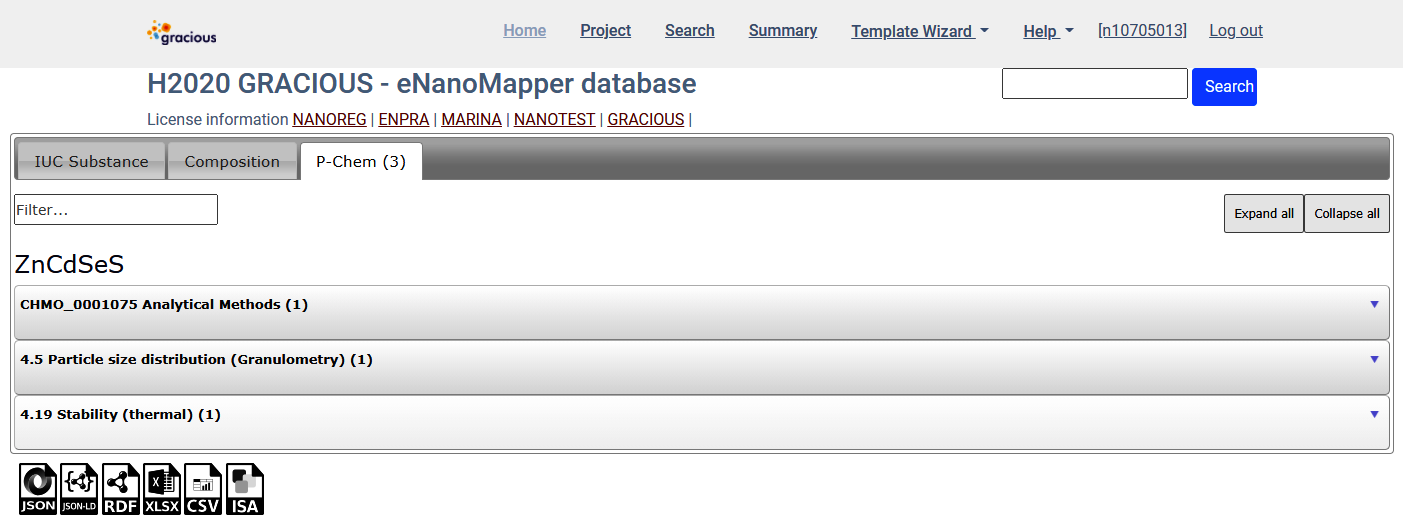
Export data for all query results - Export tab
Compact summary report
- select
Compact data reportfrom the dropdown menu, click onXLSXicon and finally click onDownload filtered entries as XLSX
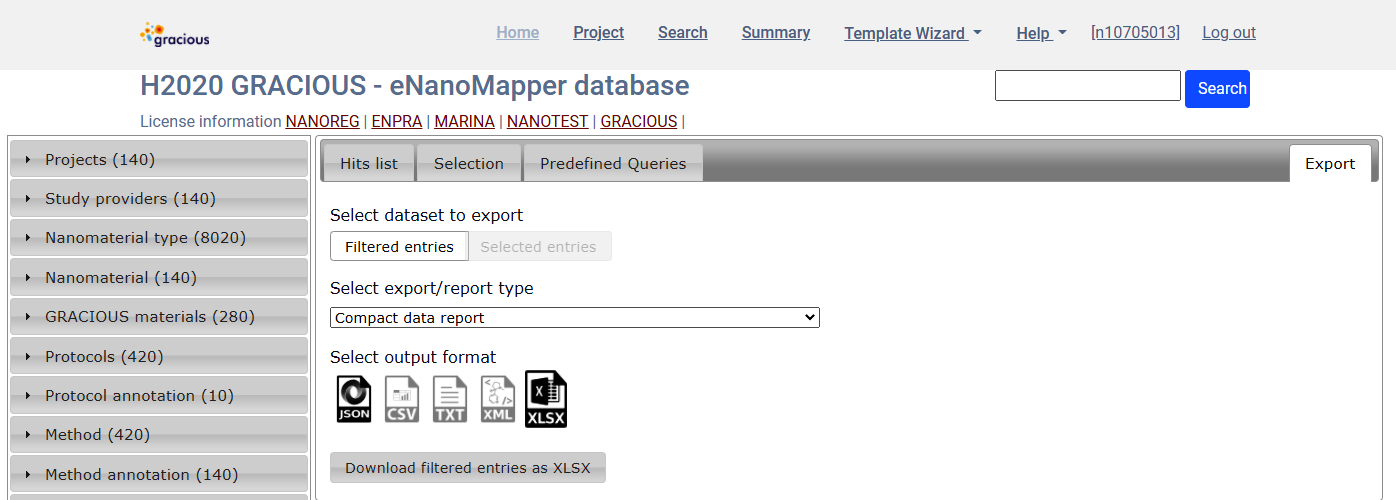
The report is an Excel file
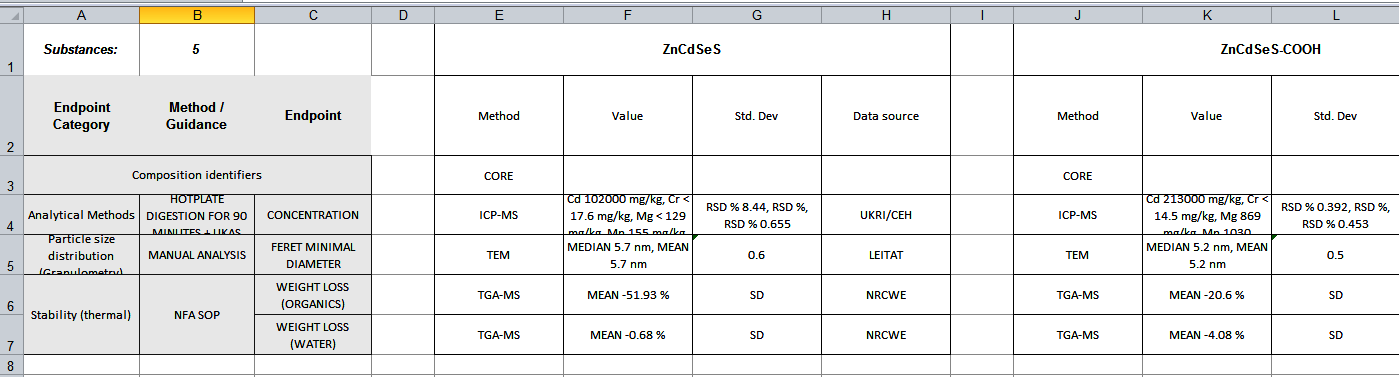
- Try exporting the same data into different report layouts (select from the dropdown menu).
Data export
Select the TXT icon and click on Download filtered entries as TSV
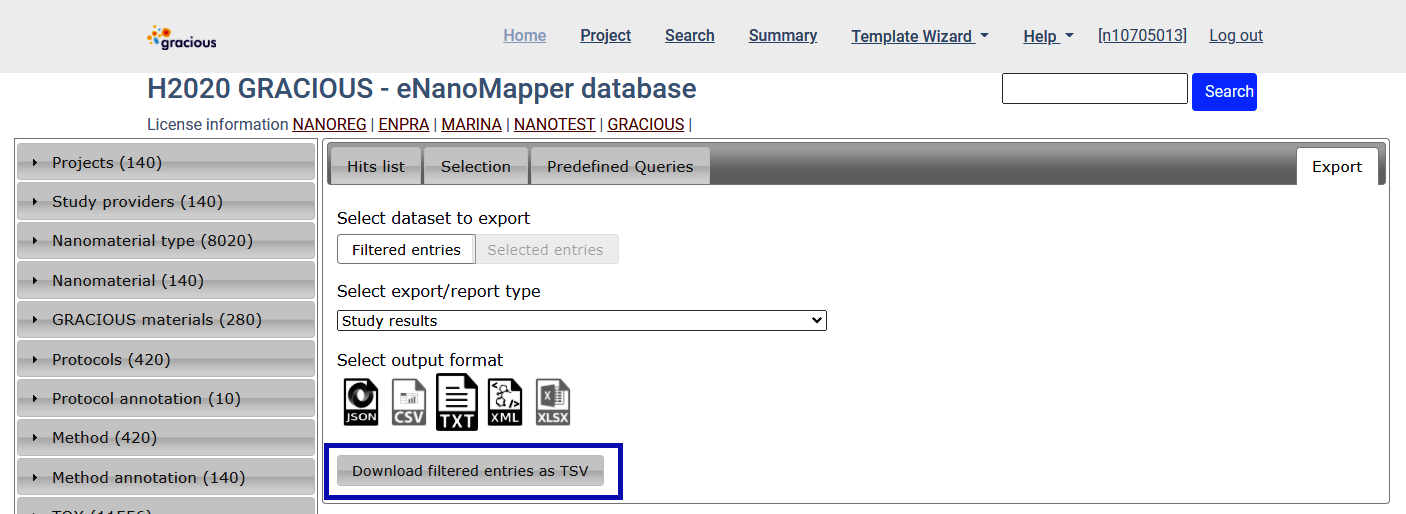
The result is a spreadsheet (.txt , tab separated text file)
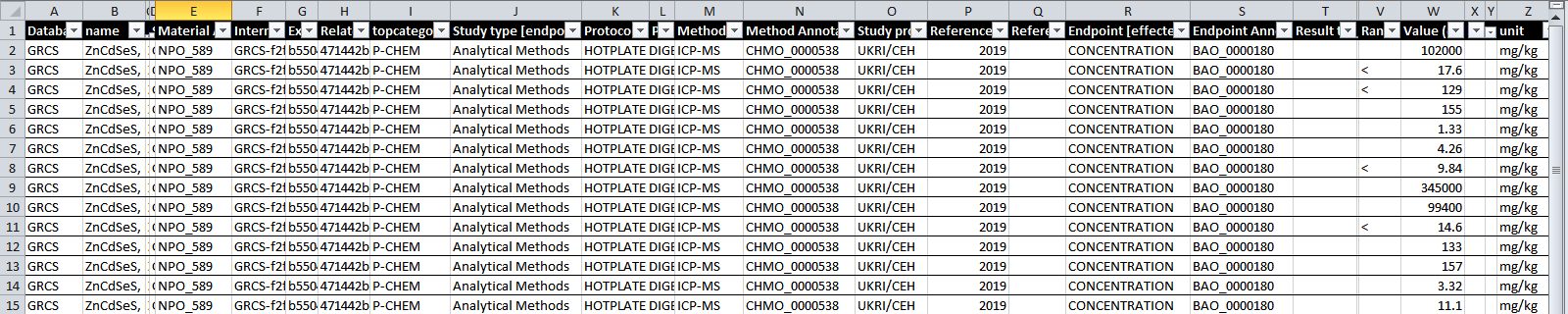
Predefined queries
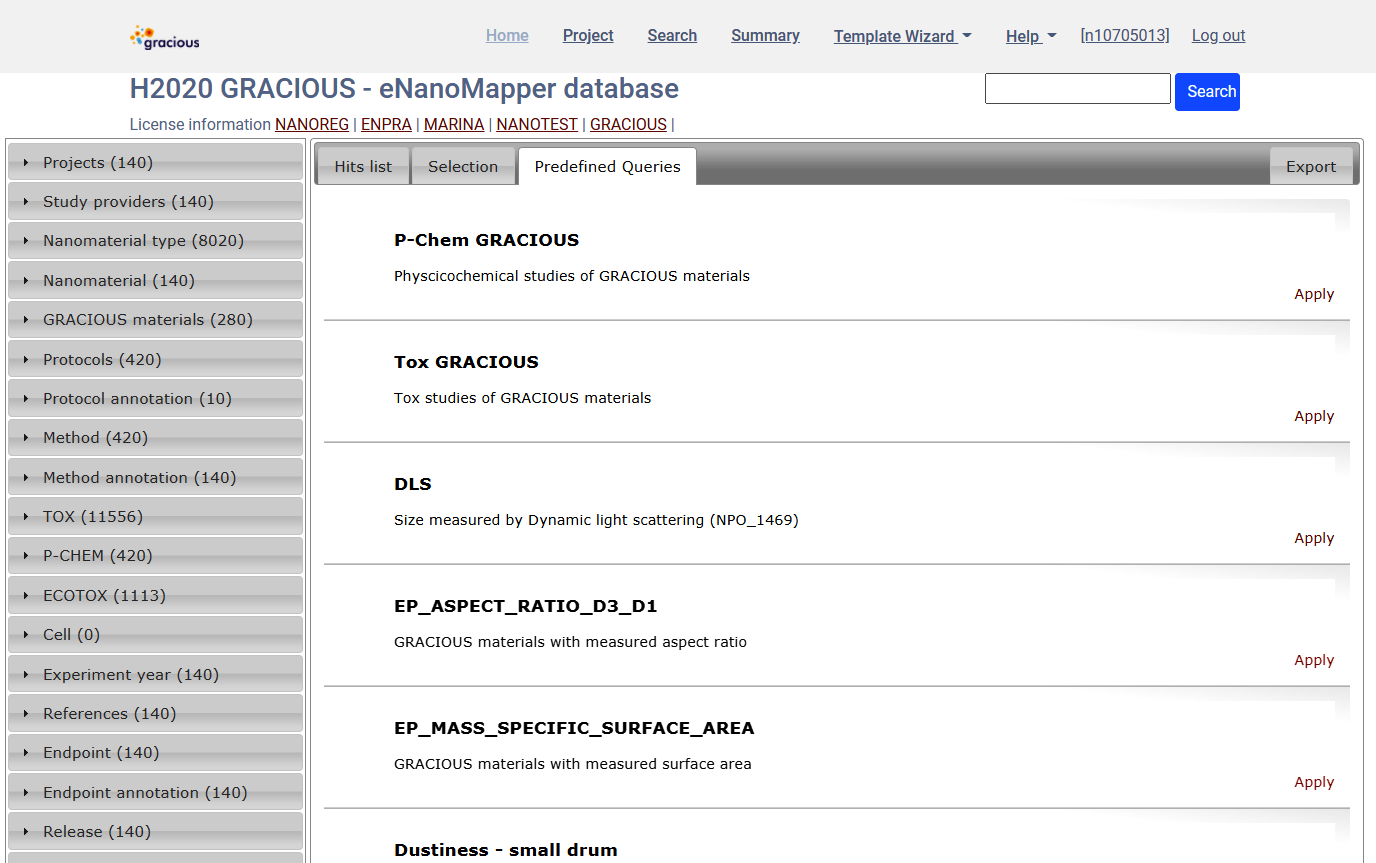
Select EP_MASS_SPECIFIC_SURFACE_AREA and click Apply. This will change the active query and will select materials with experimental data for specific surface area..
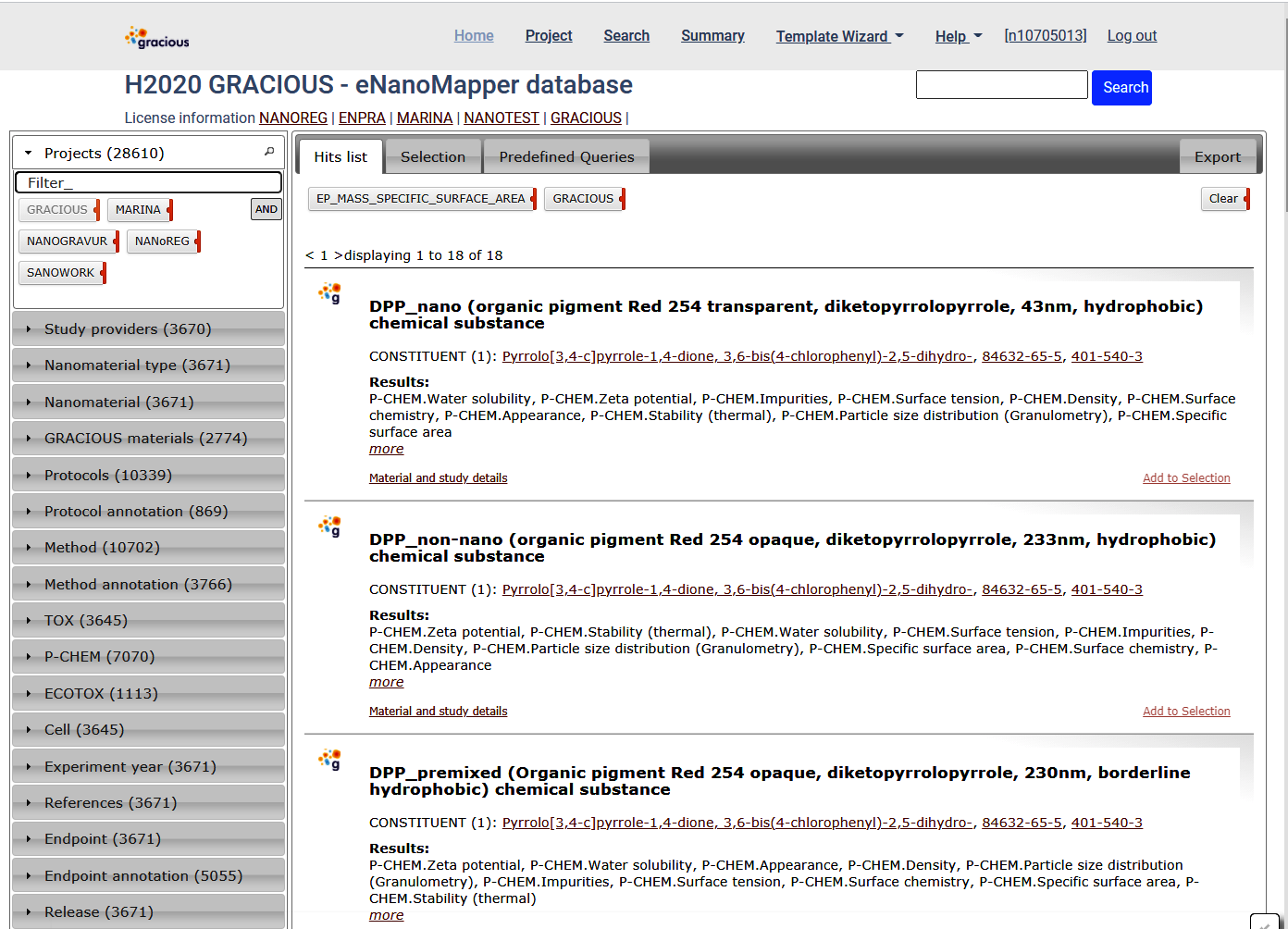
This query could be further refined using the widgets at the left, and data exported as described.
Search by ontology terms
OBI_0302736 (comet assay) is a term defined in Ontology for Biomedical Investigations (OBI) BioPortal link
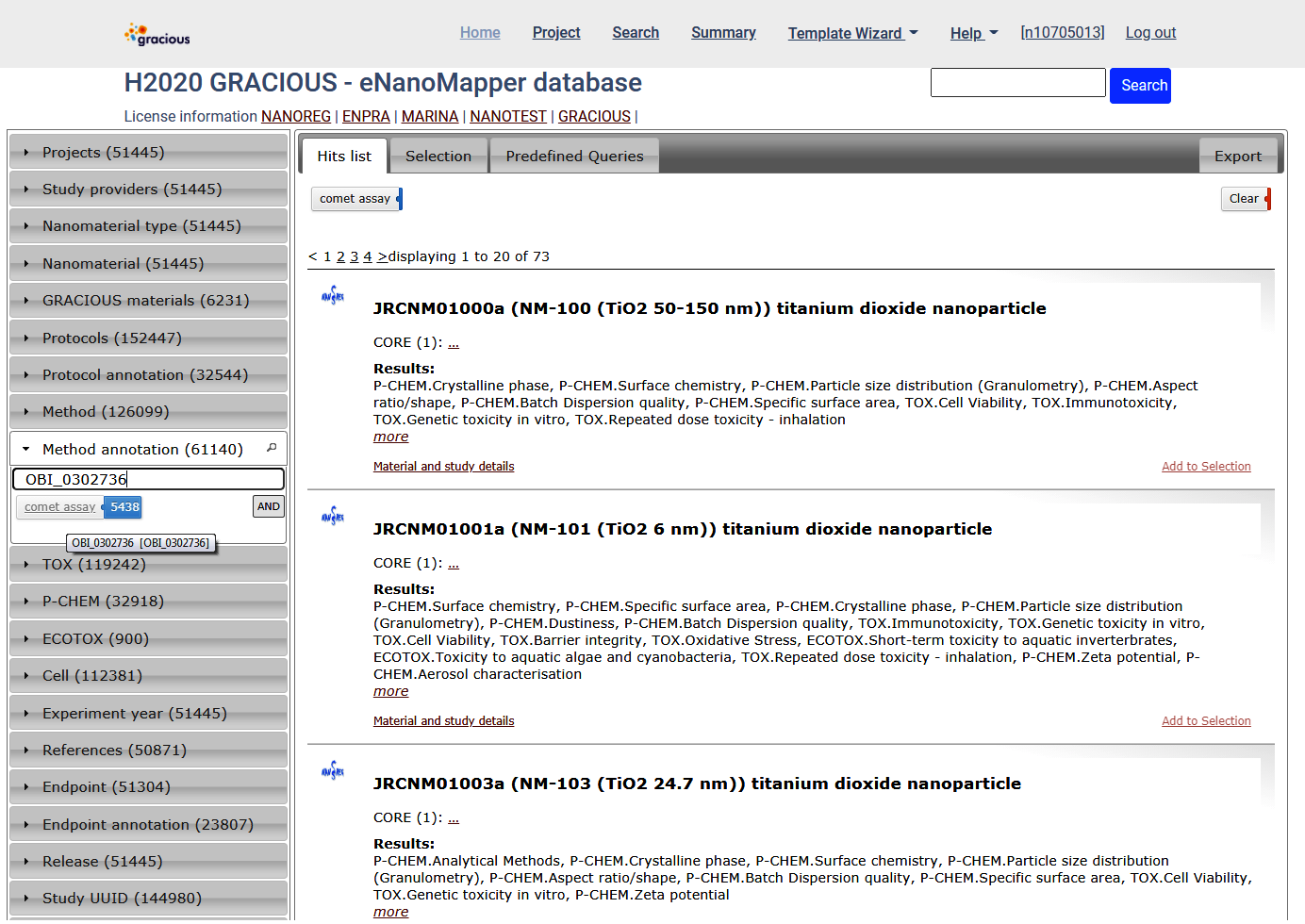
Summary
-
Various types of graphical summaries are available at the Dashboard project page
-
Download as image (SVG, PNG) or Excel files
Programmatic access (Application Programming Interface - API)
API documentation
- Online interactive API documentation with standard OpenAPI3 format
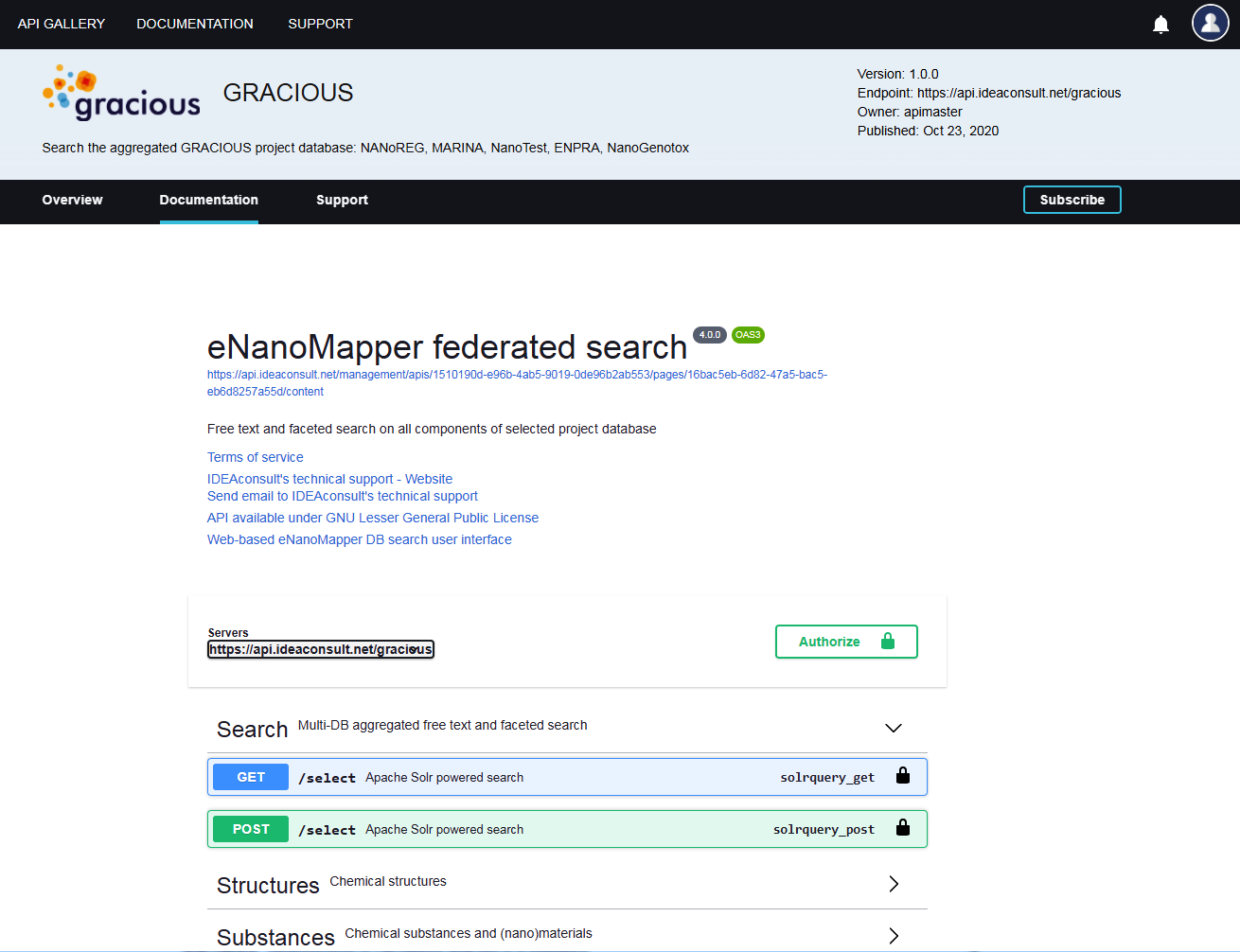
eNanoMapper client libraries
- Python : https://pypi.org/project/pynanomapper/
- R : https://github.com/enanomapper/renm
- JavaScript : https://github.com/enanomapper/ambit.js/
- JavaScript : https://github.com/ideaconsult/jToxKit
Jupyter notebooks
https://github.com/ideaconsult/notebooks-ambit/blob/master/enanomapper/tutorial.ipynb https://github.com/ideaconsult/notebooks-ambit/blob/master/enanomapper/summary.ipynb https://github.com/ideaconsult/notebooks-ambit/blob/master/enanomapper/explore.ipynb
Summary charts generated with explore.ipynb
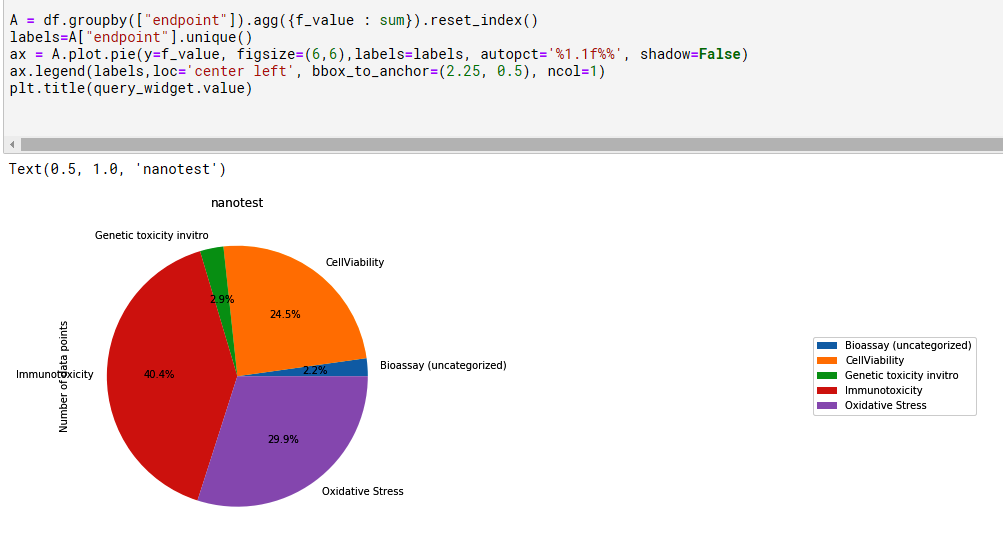
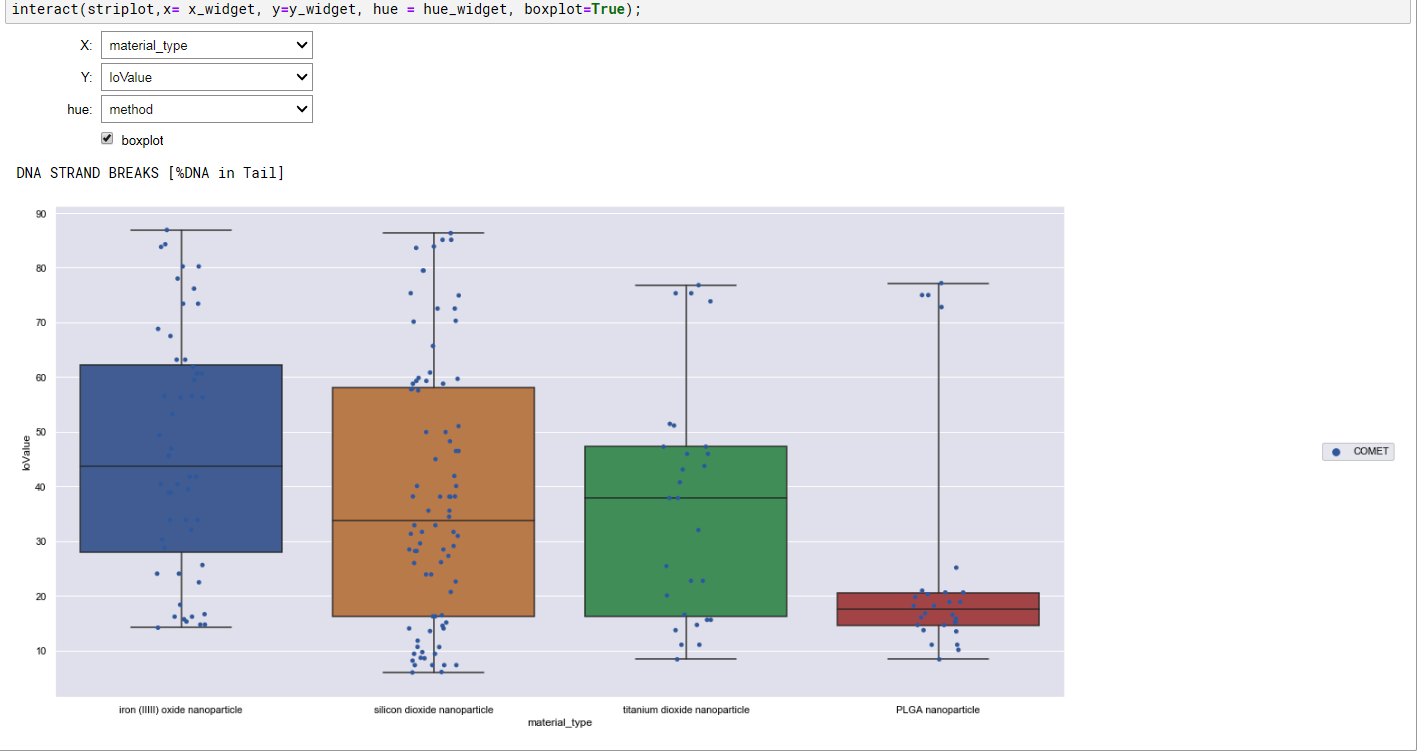
Data entry
The data entry for eNanoMapper and the Template Wizard are described in this tutorial.
