The eNanoMapper data model
represents the nanomaterials as chemical substances
1.eNanoMapper/AMBIT data model overview
The eNanoMapper data model represents NMs as chemical substances and links experimental data to the substances and not to the individual components.
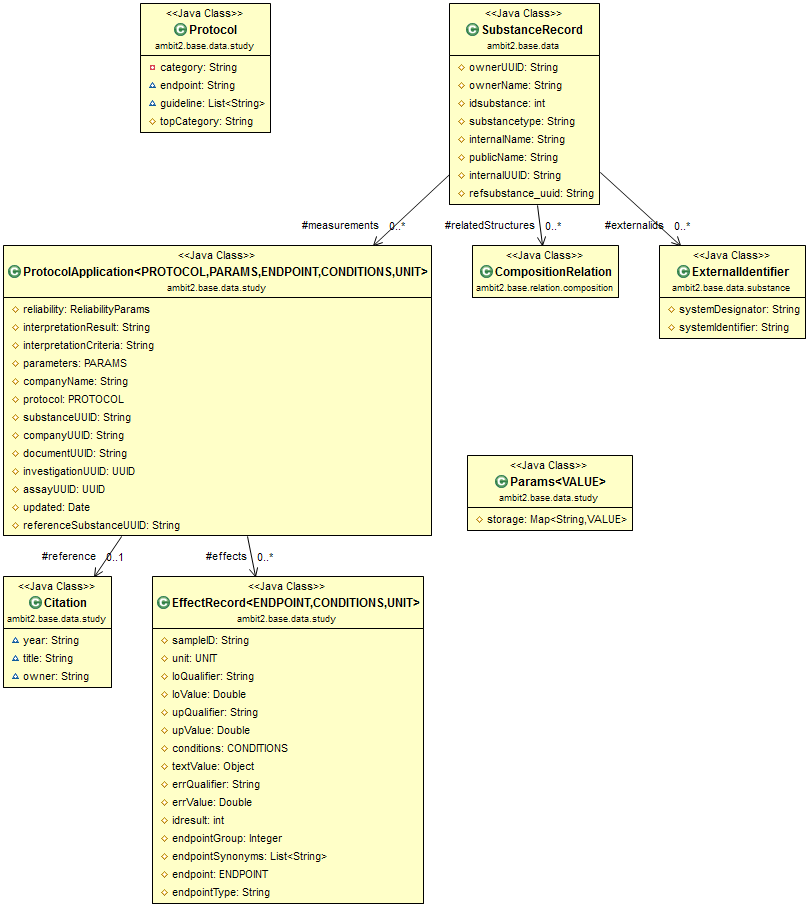 The high level components of the model such as Substance, Protocol Application, Investigation and Assay have automatically generated UUIDs.
The high level components of the model such as Substance, Protocol Application, Investigation and Assay have automatically generated UUIDs.
More information at NMDmdataParser GitHub repository
2.Representation of chemical substances and nanomaterials
The implementation of substance support in Ambit is inspired by the data models of OECD HT, CODATA, BioAssay ontology and ISA-TAB data model. The substance is represented by the SubstanceRecord class. A substance can be serialized in multiple formats when retrieved via API.Example query
curl -X GET "https://api.ideaconsult.net/nanoreg1/enm/nanoreg1/substance?search=NM-100&pagesize=10" -H "accept: application/json"
Response
{
"substance": [{
"URI": "https://apps.ideaconsult.net/nanoreg1/substance/NNRG-3c52517e-f126-4fdd-1cda-2434534e46b8",
"ownerUUID": "NNRG-04022171-cf37-b07b-b933-a6b8d57883ed",
"ownerName": "NANoREG",
"i5uuid": "NNRG-3c52517e-f126-4fdd-1cda-2434534e46b8",
"name": "NM-100 (TiO2 110 nm)",
"publicname": "JRCNM01000a",
"substanceType": "NPO_1486",
"externalIdentifiers": [{
{
"type": "NANoREG material",
"id": "Core material"
},
{
"type": "NANoREG supplier",
"id": "Fraunhofer,JRC - IHCP,IIT"
},
{
"type": "NM code",
"id": "NM-100"
}
]
},
{
"URI": "https://apps.ideaconsult.net/nanoreg1/substance/NRGF-1809fa0c-6785-3caa-989c-22cb16ec3100",
"ownerUUID": "NRGF-04022171-cf37-307b-b933-a6b8d57883ed",
"ownerName": "NANoREG",
"i5uuid": "NRGF-1809fa0c-6785-3caa-989c-22cb16ec3100",
"name": "NM-100 TiO2 50-150 nm",
"publicname": "JRCNM01000a",
"substanceType": "CHEBI_51050",
"externalIdentifiers": [{
"type": "JRC ID",
"id": "JRCNM01000a"
},
{
"type": "NANoREG supplier",
"id": "JRC"
},
{
"type": "NM code",
"id": "NM-100"
}
]
}
],
"records": 2
}
3.Substance identifiers
A substance within enanoMapper database instance (e.g. NANoREG) is assigned unique UUID (`i5uuid`), usually autogenerated on import from substance name and composition.
This is the unique identifier which can be used to retrieve substance details from the AMBIT API /enm/{db}/substance/{uuid} , composition and all study information /enm/{db}/substance/{uuid}/study.
The substance carries two name fields - substance name and publicname
The substance name is unique within the database instance , and typically is a project-assigned name of the material. It is typical that projects assigns internal names to materials, and this is what substance name is for.
Where possible, the publicname contains a widely accepted unique identifier (e.g. JRCNM reference identifier or else ), which could be used to match materials across projects.
The substanceType indicates the material type (i.e. Titanium dioxide nanoparticle) using the eNanoMapper ontology terms (e.g. CHEBI_51050).
A substance may contain details on composition. The substance composition is not just a simple list of constituents but complex relations between the constituents may exist. Each constituent may have its own properties (typically nominal physichem properties) which are distinguished from the substance properties. Each constituent is assigned a role in the composition such as main constituent, impurity or additive or for cases of nanomaterials the roles can be core, coating 1, coating2 etc.
Arbitrary number of external identifiers can be assigned (e.g. ERM ,identifiers used in other databases).
4.Representation of experimental data
Substance physicochemical and biological properties are described by means of rich experimental data and supporting metadata used for the interpretation of the experimental values and is represented in a multi-layer fashion (see also ISA-TAB data model). The event of applying a test or experimental protocol to a substance/nanomaterial is described by aprotocol application entity.
Each protocol application consists of a set of measurements for a defined endpoint under given conditions or experimental factors.
The measurement result can be a numeric value with or without uncertainty specified, an interval, a string value, or a link to a raw data file (e.g., a microscopy image). Particular “protocol application” is distinguished by a dynamic list of specific parameters. The measurement conditions are another dynamic list of parameters put in a different level of the data model. The data for particular NM may contain many “protocol applications”. The protocol applications which are related can be grouped to form an “Investigation” entity.
Also several different nanomaterials/substances can which have
protocol application applied could be grouped via the Assay entity.
6.Server implementation
The eNanoMapper data model is implemented in an open source software, AMBIT, hence all implementation details (including database structure) can be examined.
7.Client libraries
- jToxKit JavaScript client library
- pynanomapper Python client library
8.References
- Kochev, Nikolay, Nina Jeliazkova, and Ivanka Tsakovska. 2019. “CHAPTER 3. Chemoinformatics Representation of Chemical Structures – A Milestone for Successful Big Data Modelling in Predictive Toxicology.” In Big Data in Predictive Toxicology, edited by Daniel Neagu and Andrea Richarz, 69–107. RSC Publishing. doi:10.1039/9781782623656-00069.
- Jeliazkova, Nina, Charalampos Chomenidis, Philip Doganis, Bengt Fadeel, Roland Grafström, Barry Hardy, Janna Hastings, et al. 2015. “The eNanoMapper Database for Nanomaterial Safety Information.” Beilstein Journal of Nanotechnology 6 (July): 1609–34. doi:10.3762/bjnano.6.165.
- Jeliazkova, Nina, Philip Doganis, Bengt Fadeel, Roland Grafstrom, Janna Hastings, Vedrin Jeliazkov, Pekka Kohonen, et al. 2014. “The First eNanoMapper Prototype: A Substance Database to Support Safe-by-Design.” In 2014 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 1–9. IEEE. doi:10.1109/BIBM.2014.6999367.
- Jeliazkova, Nina, and Vedrin Jeliazkov. 2011. “AMBIT RESTful Web Services: An Implementation of the OpenTox Application Programming Interface.” Journal of Cheminformatics 3 (1): 18. doi:10.1186/1758-2946-3-18
